The Slovak Centre of Scientific and Technical Information (CVTI SR) is building the Slovak EOSC Node to integrate national research resources into the EOSC Federation. Through its KOMIS interoperable platform, the Node connects Slovak research organisations, universities, and data repositories, including the Research Data Objects Repository (SVD).
CVTI SR also leverages substantial infrastructure, including the Datacentre for Research and Development, alongside specialised bioinformatics environments and HPC resources interconnected with Slovak research stakeholders. Key contributions include FAIR-compliant data management and cross-border research collaboration capabilities in the form of cross-node scientific use cases.
CVTI SR’s combined role as National EOSC SK Secretariat and National Office for Open Science positions the Slovak EOSC Node as a bridge between Slovak research infrastructures and the broader European research ecosystem, addressing data fragmentation while enhancing interoperability and accessibility for Slovak and European researchers.
Key objective: To enhance Slovakia’s research capabilities, contribute Slovak resources and expertise, address the fragmentation of Slovak research data, and improve data accessibility. The initial technical objective involves integrating KOMIS subsystem SVD while ensuring compatibility with FAIR principles and other EOSC Federation standards. Subsequent goals include achieving integrations of the Datacentre of CVTI SR and establishing interconnections with HPC-related services of Slovak research stakeholders.
Science areas: Multidisciplinary—the initial build-up phase will focus particularly on bioinformatics and biomedical research domains, such as antimicrobial resistance research and AI for medical diagnostics.
FAIR data
FAIR-compliant repositories are integrated via KOMIS (mainly subsystem SVD – Research data management system, secondary SK CRIS, SCIDAP, etc.), and persistent identifiers (DOI, ORCID, ROR) are implemented. Metadata harmonisation ensures interoperability across Slovak and European resources, with OAI-PMH compatibility ensured.
Scientific use cases
The Slovakian EOSC Node’s scientific impact extends beyond individual use cases by establishing replicable patterns, demonstrating practical EOSC interoperability, and providing a model for smaller countries to progressively engage with EOSC while building local capacity. These initiatives collectively enhance European research capabilities while maintaining data sovereignty and enabling effective cross-border scientific collaboration.
Federated analysis of pathogen genomes
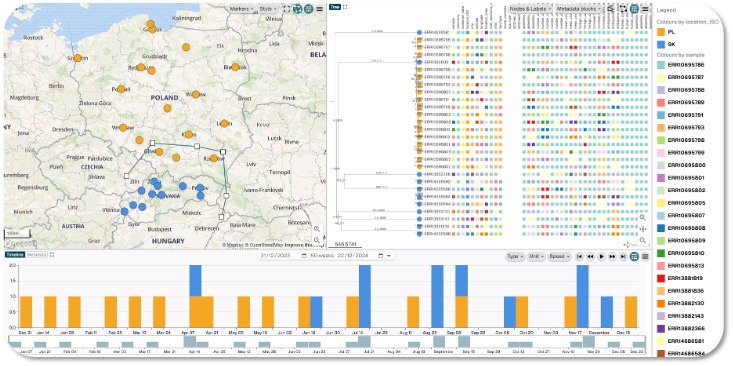
The federated analysis of pathogen genomes science case outlines a federated, cross‑border capability for timely analysis of pathogen genomes that brings computation to the data instead of copying sensitive datasets across institutions.
The objective is to shorten time‑to‑insight for outbreak detection, source attribution, and antimicrobial‑resistance (AMR) surveillance while preserving data sovereignty and meeting European legal and ethical requirements. Experience from COVID‑19 showed that sequencing at scale can transform public‑health decision‑making. Operationally, the effort starts with two neighbouring nodes of the EOSC Federation—the Slovakian national node providing workflows, datasets, computational infrastructure and domain expertise, and the Polish national node (via Poland’s National Science Centre (NCN) and a scientific repository service) supplying key technical support and their own datasets. Their geographical proximity make the two EOSC Nodes an ideal pair for a cross-border pilot. The approach demonstrates how to establish a federation of trusted sites, run harmonized workflows locally, and share only the minimum results needed for action.
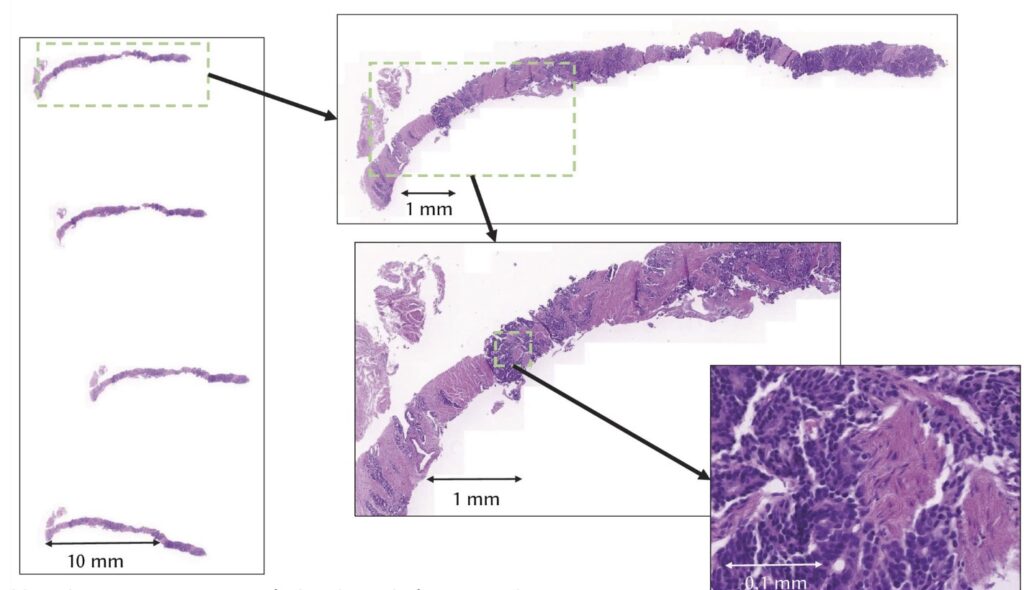
Multi-centric validation of AI models for prostate-cancer screening
Advances in digital pathology are transforming cancer diagnosis by enabling high-resolution imaging of tissue samples and the application of artificial intelligence (AI) for clinical decision support.
Prostate cancer, one of the most prevalent malignancies among men worldwide, represents a critical case for early and accurate diagnosis, as survival rates are strongly tied to timely detection and treatment. The objective of the multi-centric validation of AI models for prostate-cancer screening (MCVAL) use case, coordinated by the BBMRI-ERIC EOSC Node, is to create a secure environment in which AI models for prostate cancer screening can be validated using data from different hospitals. Rather than building new diagnostic systems from scratch, the project focuses on testing an existing model trained on whole-slide images and assessing how well it performs when applied to data processed elsewhere.