The Life Sciences Connect EOSC Node federates data, tools and services from major European life science infrastructures, providing access to biomolecular, biomedical, imaging and structural biology resources.
It enables FAIR-compliant integration of omics, imaging, clinical and experimental model data, supporting cutting-edge research and innovation. In addition, it delivers virtual research environments, workflows and secure access frameworks in line with GDPR and ethical guidelines.
Key objective: To augment the biodata expertise of the EOSC Federation, ensuring interoperable, FAIR, secure and ethically compliant access to life science data, services and computational resources across Europe.
Science areas: Life sciences, including genomics, proteomics, metabolomics, imaging, neuroscience, clinical research, systems biology, and bioinformatics.
FAIR data
The Life Sciences Connect EOSC Node federates FAIR omics, imaging, structural and clinical data repositories, along with FAIR compliant software, workflows and training to accelerate discovery, support personalised medicine and strengthen European competitiveness in life sciences.
Imaging data workflows on Galaxy
This science case on imaging data workflows on Galaxy demonstrates how a federated, open-access computational platform can transform the way imaging data are processed, shared, and reused across diverse scientific domains.
The use case leverages the Galaxy platform to integrate data from a wide range of imaging-based research fields—such as life sciences (microscopy), astrophysics (telescope data), climate science (satellite data), and marine science (underwater imagery)—into a unified analysis environment. The goal is to demonstrate that Galaxy can be integrated into the EOSC Federation’s common infrastructure to serve many disciplines simultaneously, allowing researchers to share workflows, reuse methods, and access powerful computational tools without needing specialised technical expertise.
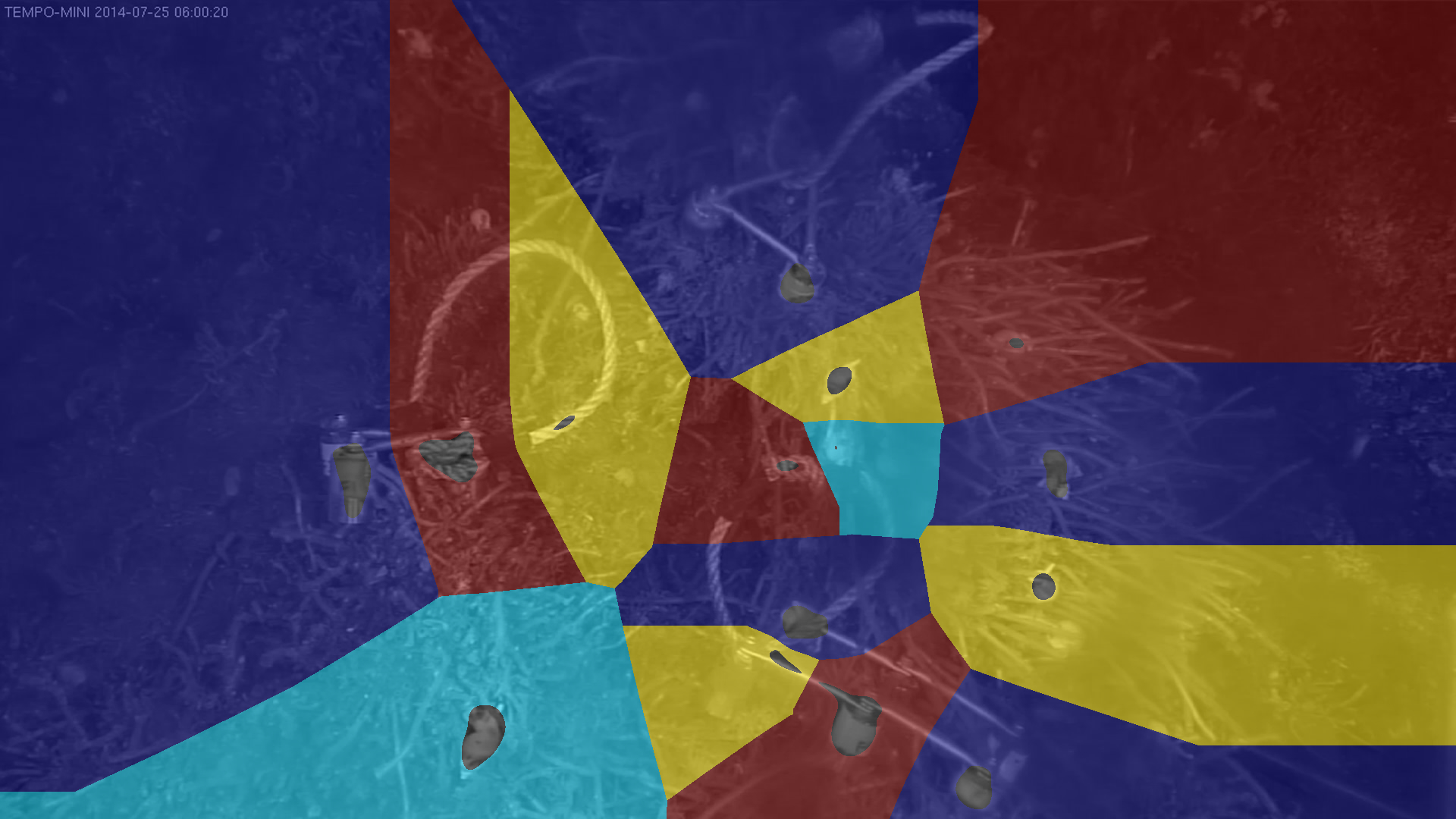

Biological sequestration of carbon in the ocean
The scientific use case addresses the ocean’s critical role in mitigating climate change through biological carbon sequestration, a process by which marine microorganisms capture atmospheric carbon dioxide and store it in the deep ocean for centuries.
Despite its global importance for climate modelling, this mechanism remains poorly represented in current models, which still rely on simplified representations of marine ecosystems. The use case seeks to fill this knowledge gap by integrating genomic, environmental, and modelling data within a unified, open, and interoperable framework provided by the EOSC Federation.
Other use cases
- Cross-domain interactive image data analysis with BAND virtual desktop: Enable human-in-the-loop analysis workflows for large scale remote datasets and support training
- Federated data storage of structural biology data: Federated access to distributed data and orchestration of data flow from instruments to repositories